Sometimes, you need to find and group together the replicated records in a spreadsheet. There are several different ways to identify duplicated records (see this tutorial for a good one), but what I wanted was something a bit more fancy. I wanted not just the duplicates, but each original record as well. Furthermore, I wanted any replicates (original + duplicates) grouped together in neat little sets in the spreadsheet.
Here is how I did it using the Unix command line.
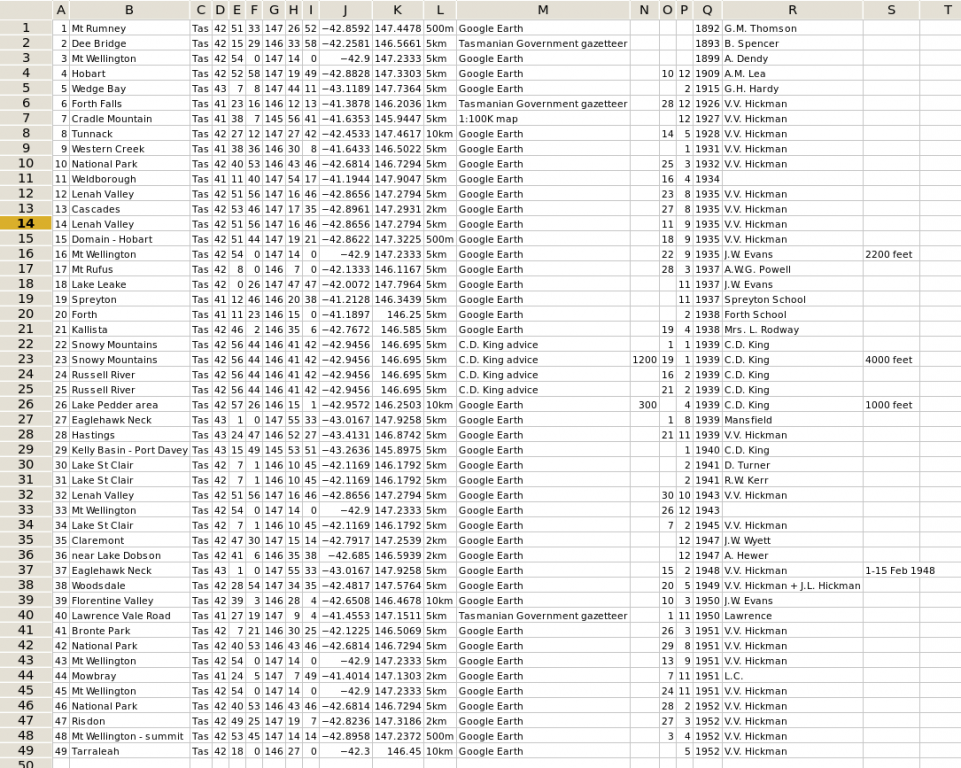
My starting point was the table of records shown in the first screenshot, except that the screenshot shows only the first 49 records out of more than 8000. Each record has a unique identifying number in the first column. What I was looking for were records with the same latitude and longitude, as listed in columns J and K.
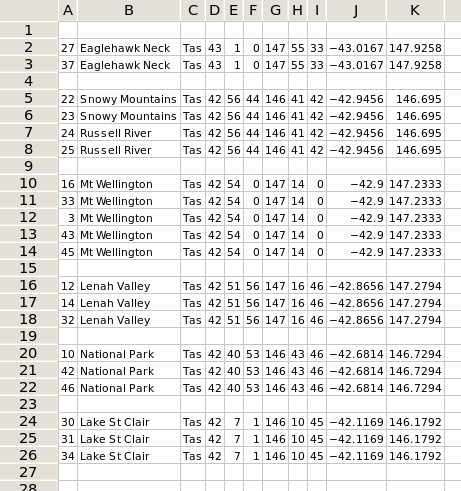
I wanted what's shown in the second screenshot: records with the same latitude and longitude grouped into sets, with the unique identifying numbers still attached in case I needed to check other details of the records.
If you work a lot with spreadsheets, you're probably thinking that a fiddly job like this would take quite a while. It actually took less than one minute to do the whole set of 8000+ records.
No magic needed
The secret was to change the spreadsheet into a text file. I could then use simple but powerful text-manipulating tools on the command line to do the job I wanted, before returning the finished text to the spreadsheet program. Here's what I did:
1. I saved the spreadsheet as a tab-separated text file. In my case, it was called file1.txt (no kidding!).
2. I opened a terminal, changed to the directory containing file1.txt, and entered the following command:
cut -f1-11 file1.txt | sort -t$'\t' -k10n,10 -k11n,11 | rev | uniq -w16 --all-repeated=prepend | rev > file2.txt
3. I opened file2.txt in a spreadsheet program. Done.
How it works
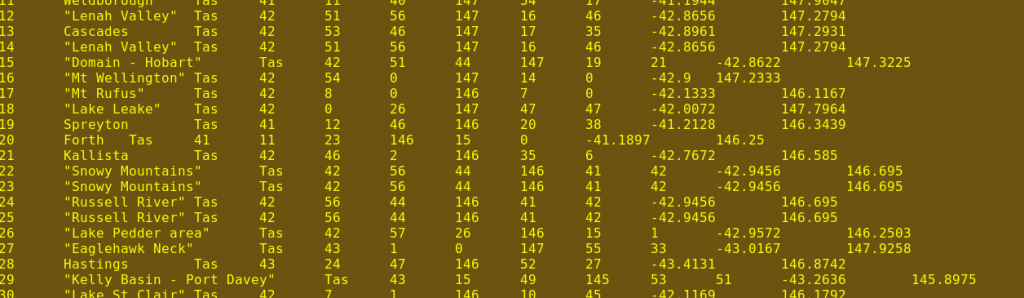
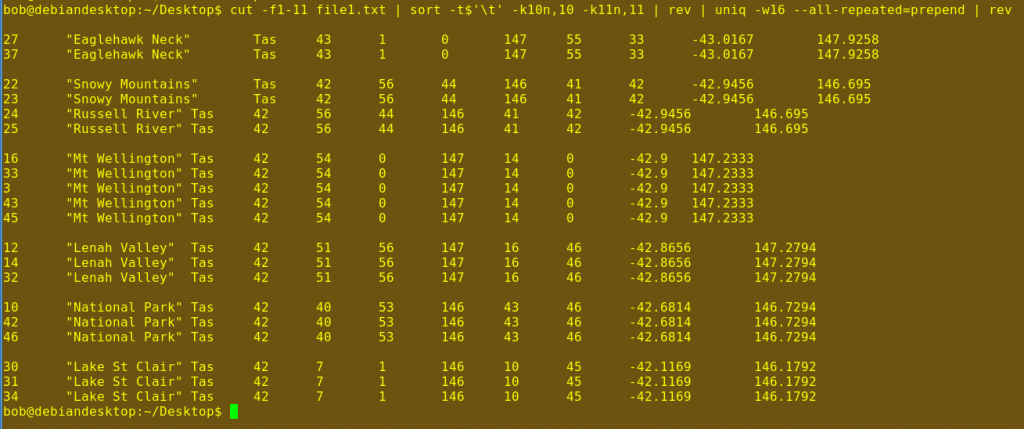
The first command, cut, is told with -f1-11 to trim each line in file1.txt to the first 11 columns (A-K in the first screenshot). You'll see why in a moment. The third screenshot shows a portion of the result.
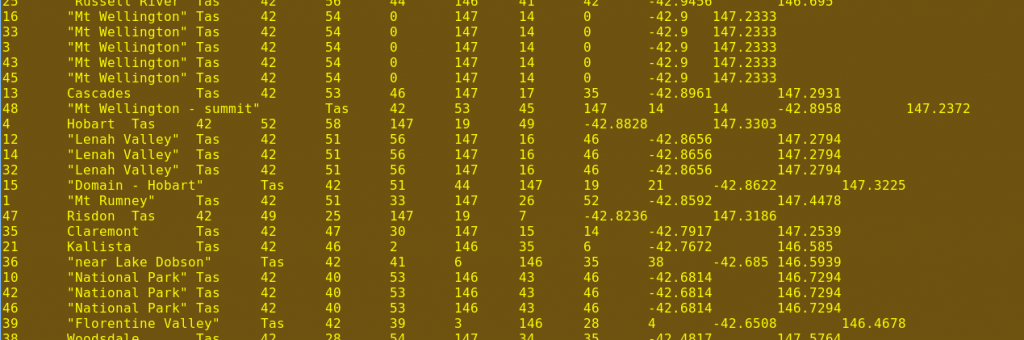
The cut file is piped (|) to the sort command, which is told with -t$'\t' that each line is broken into fields by tab spacing. This is because the $ sign means "interpret backslashes in the following string', which means that '\t' becomes the "tab" character. The command sort is also told with -k10n,10 to first sort the whole file on the contents of field 10 (column J), treating the contents as a number (n). When that's finished, sort then sorts the file again on the contents of field 11 (column K). The result is that the file is sorted by latitude and longitude with originals and duplicates in adjacent lines, as shown in the fourth screenshot.
Now a tricky bit. The sorted file is piped to the rev command, which reverses each line (fifth screenshot).
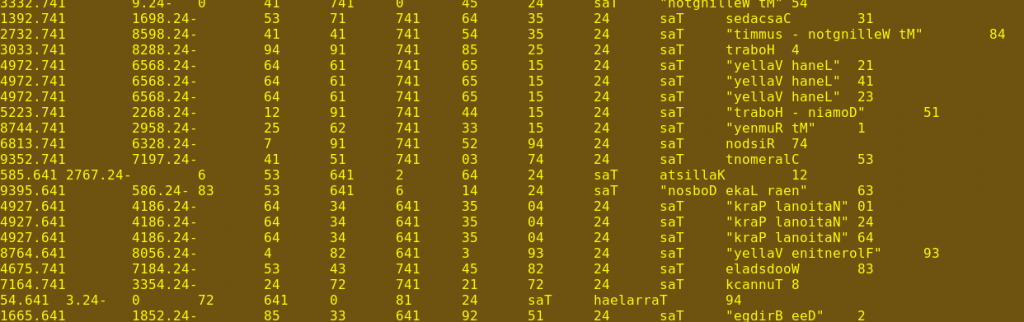
Notice in that last screenshot that in replicated records, the first 16 characters of the line are the same, even though the ends of the lines are different. This reversed file is piped to the uniq command, which is told with -w16 to look only at those first 16 characters. With --all-repeated=prepend, uniq is also told to discard all unique lines (lines without duplicates) and to separate the resulting groups with a blank line, as shown in the sixth screenshot.
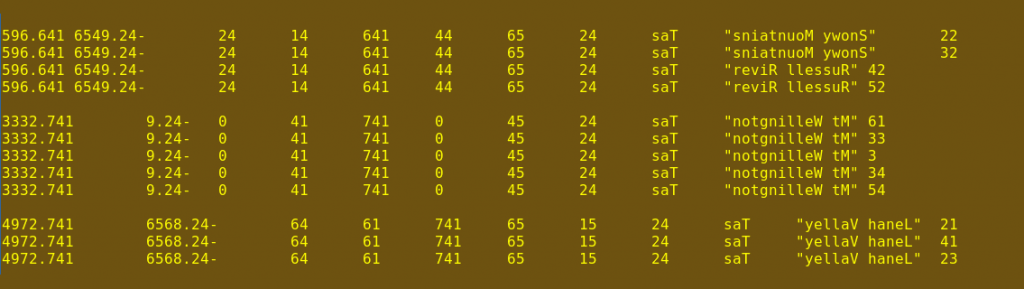
Next, the file is piped to the rev command again to get the characters back in their original order (seventh screenshot). Finally, the neatly grouped sets of replicates are sent to file2.txt, which is automatically created by the > command. The new file file2.txt is tab-separated and easily opened as a spreadsheet.
Love that command line
Another handy feature of the command-line approach is that my complicated command can be saved as a bash script – I don't have to remember it. When I want to do this job again, I just save the spreadsheet as file1.txt and run the script from a launcher. The command line really is a time saver!
For more on cut, sort and uniq, see the GNU manual for the coreutils suite of tools.